Leland McInnes
@leland_mcinnes
Followers
5,864
Following
819
Media
79
Statuses
3,864
A mathematician dabbling in the world of data science. Researcher at the Tutte Institute for Mathematics and Computing. UMAP, HDBSCAN, PyNNDescent. He / Him.
Ottawa, Ontario
Joined October 2016
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
ジェンティルドンナ
• 122735 Tweets
#Solingen
• 92556 Tweets
満塁ホームラン
• 87481 Tweets
大谷翔平
• 76911 Tweets
TO MAESTRO KHEM WITH LOVE
• 70486 Tweets
Brighton
• 66535 Tweets
#ラヴィットロック2024
• 48025 Tweets
史上最速
• 45225 Tweets
Vielfalt
• 40809 Tweets
#V最協S6
• 35877 Tweets
ML IN MACAU
• 32896 Tweets
ケンタッキー
• 29733 Tweets
Täter
• 29577 Tweets
Gabbar
• 29418 Tweets
大谷さん
• 28280 Tweets
Cemal Enginyurt Tutuklansın
• 27338 Tweets
Slogan
• 26155 Tweets
#FNTHWIN
• 26013 Tweets
Messer
• 25479 Tweets
悪役令嬢の中の人
• 24728 Tweets
グランドスラム
• 23699 Tweets
大谷選手
• 22243 Tweets
江戸川花火大会
• 19652 Tweets
Kアリーナ
• 18431 Tweets
オオタニサン
• 16940 Tweets
ALNP FANMEET IN HK
• 14985 Tweets
スーパースター
• 14206 Tweets
STRAY KIDS DOMINATE SEOUL
• 10845 Tweets
Last Seen Profiles
Understanding UMAP - an interactive introduction to the algorithm and how to us (and mis-use) it from
@_coenen
and
@adamrpearce
. A must read for anyone interested in dimension reduction.
7
232
653
UMAP 0.4 is now out! It includes a host of new features, including plotting support, better sparse data support, inverse transforms, and embedding to non-euclidean manifolds.
pip install umap-learn
See this thread for some of the new features:
5
178
584
Really enjoying the
#mlprague
conference. Slides for my talk on topological approaches to unsupervised learning problems can be found here:
6
66
260
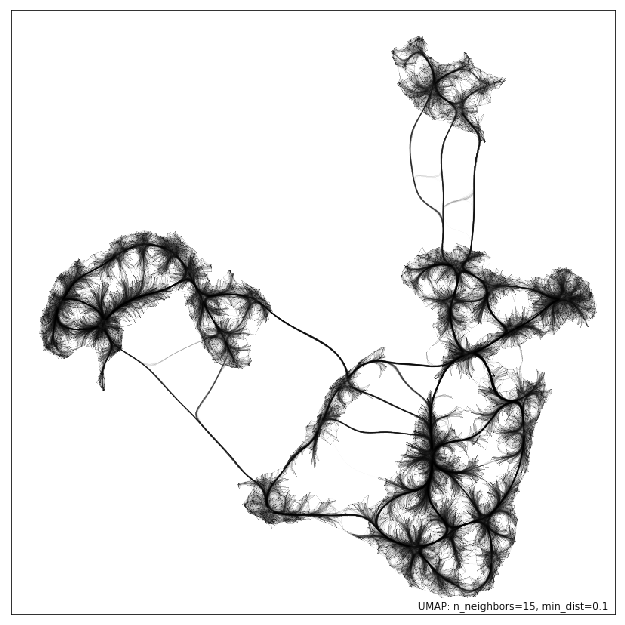
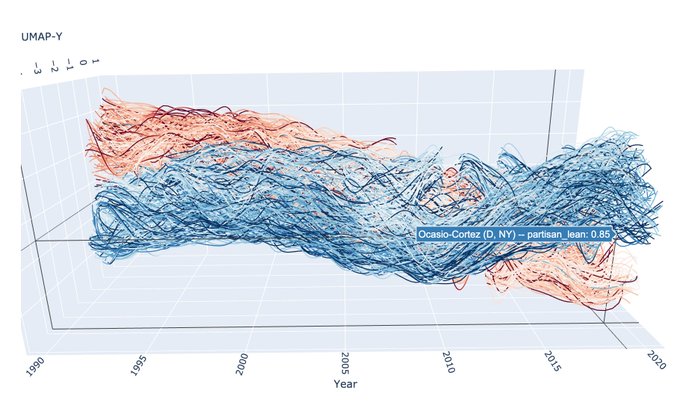
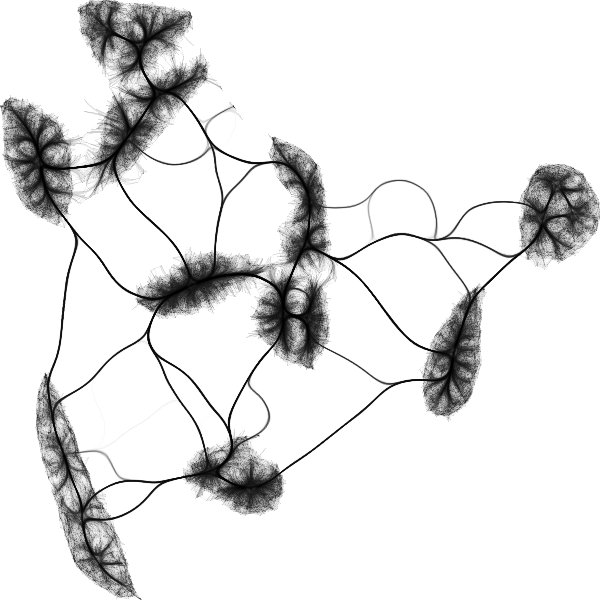
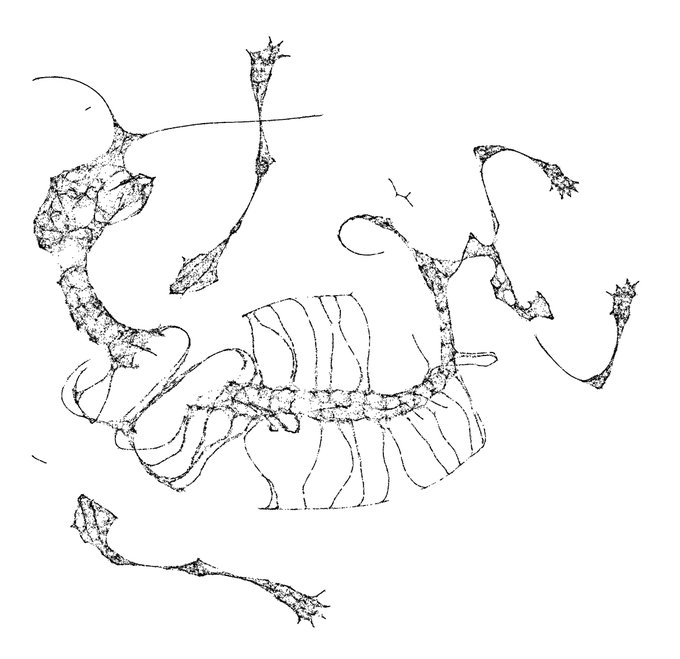
I just started playing around with
@datashader
edge bundling for visualizing graphs associated to UMAP embeddings. Here's one for MNIST:
7
29
182
This is some amazing work from
@tim_sainburg
. Some major takeaways:
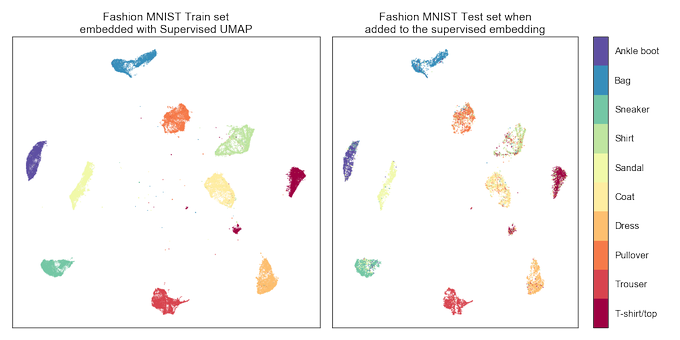
- lightning fast transform/inverse_transform operations (comparable to PCA if you have a GPU);
- semi-supervised classification: 97.8% accuracy on MNIST with only 4 labelled items per class!
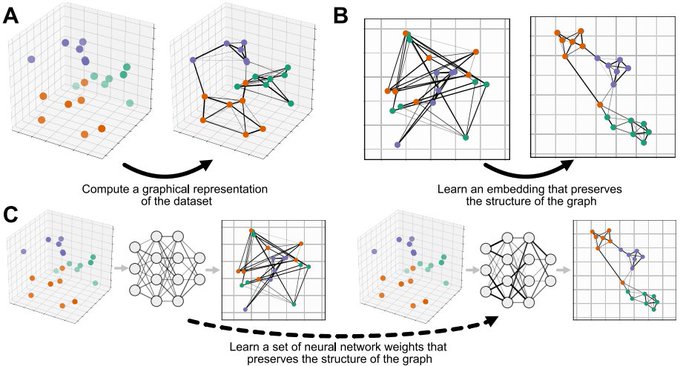
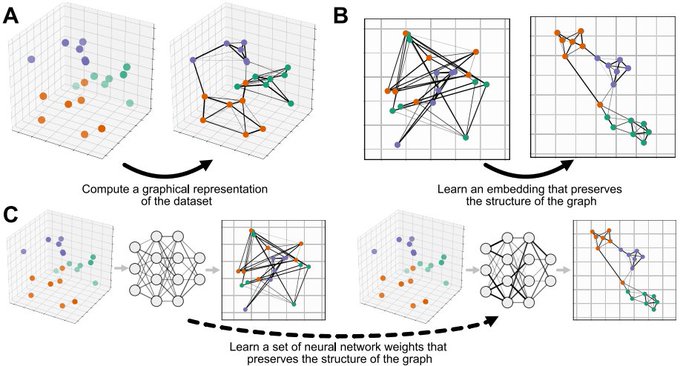
New paper "Parametric UMAP: learning embeddings with deep neural networks for representation and semi-supervised learning" with
@leland_mcinnes
and
@TqGentner
! 1/
4
61
294
1
38
154
I just gave a talk at
#PyDataNYC
on dimension reduction -- focussing on core intuitions and unifying concepts. You can find the slides for it here:
5
52
129
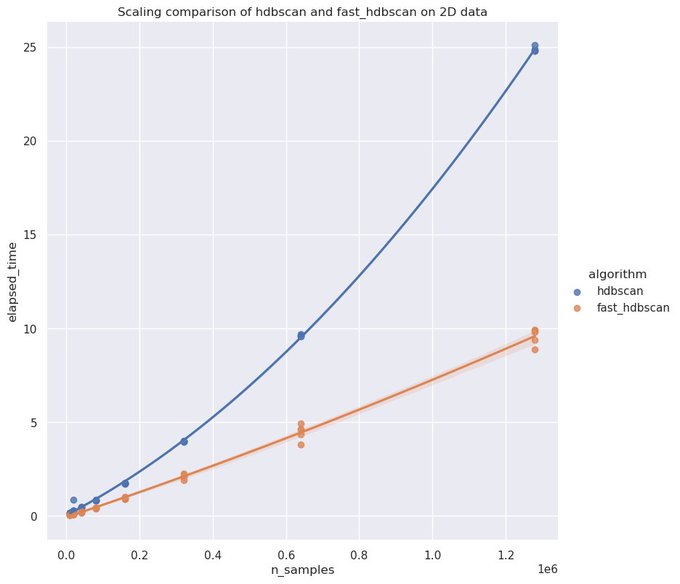
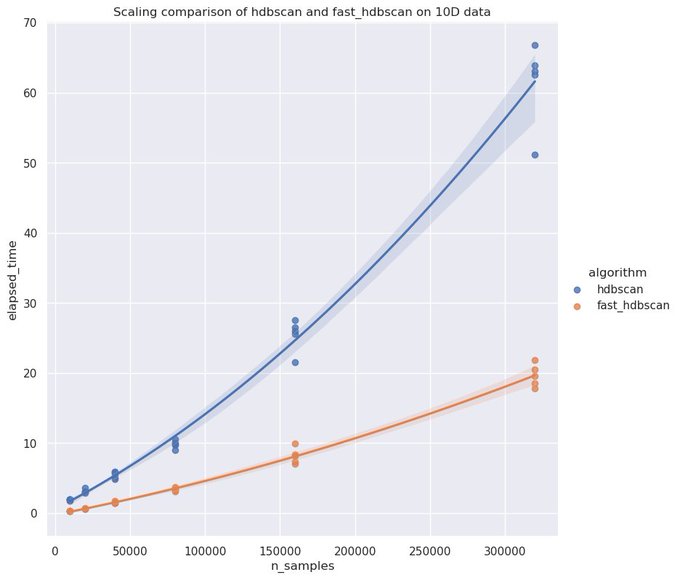
If you have GPU resources handy the new HDBSCAN implementation in
@RAPIDSai
cuML is amazingly fast. You can get to millions of points clustered in only a few minutes!
3
20
112
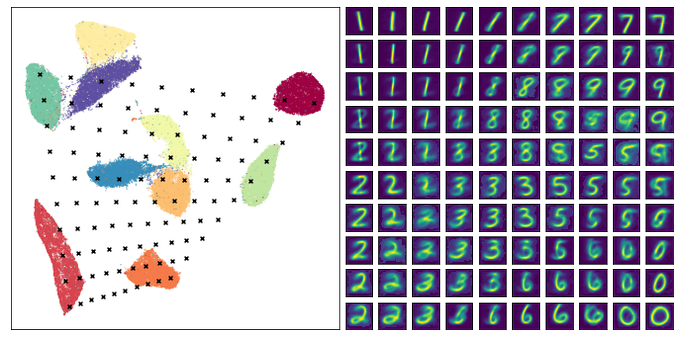
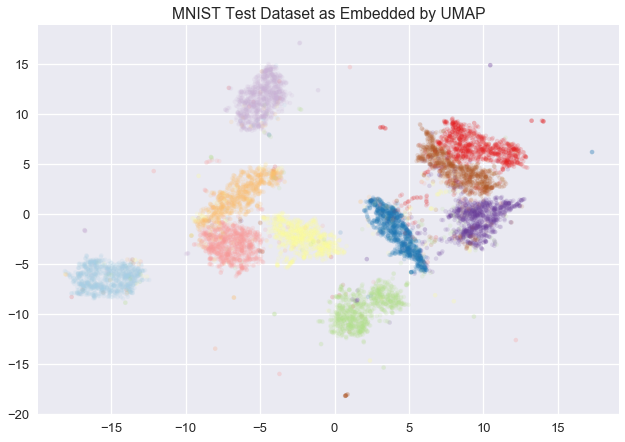
If you want to spend some time exploring a UMAP embedding of images (like MNIST)
@GrantCuster
put together a nice tool:
2
37
104
My
#scipy2021
talk on PyNNDescent, a library for fast approximate nearest neighbour search is now available:
5
23
97
A great example of what UMAP is for: look at your data and realise it wasn't what you thought -- and then use it to ask better questions about your data before proceeding with fancier ML tools.
0
21
95
My talk on topological data analysis at ML Prague is already online! It provides a brief whirlwind tour of why topological methods matter for unsupervised learning problems.
#mlprague
0
35
92
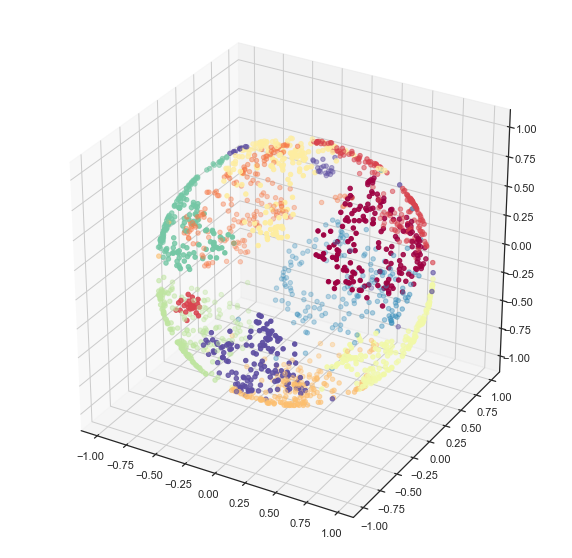
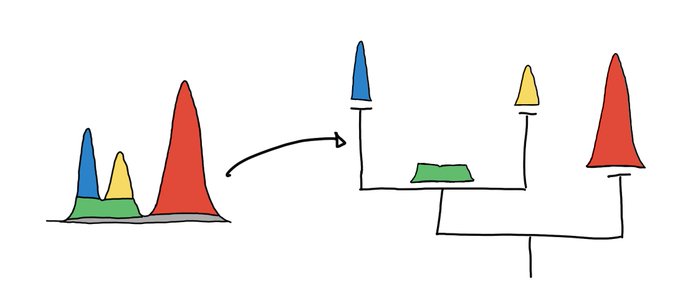
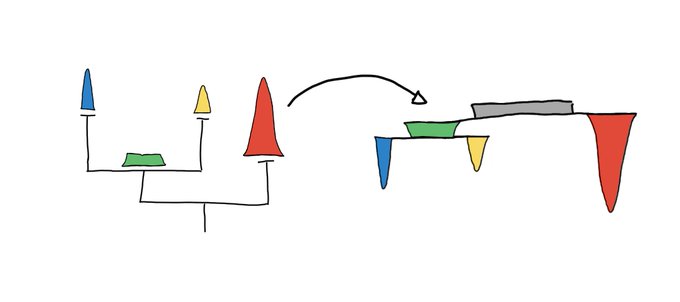
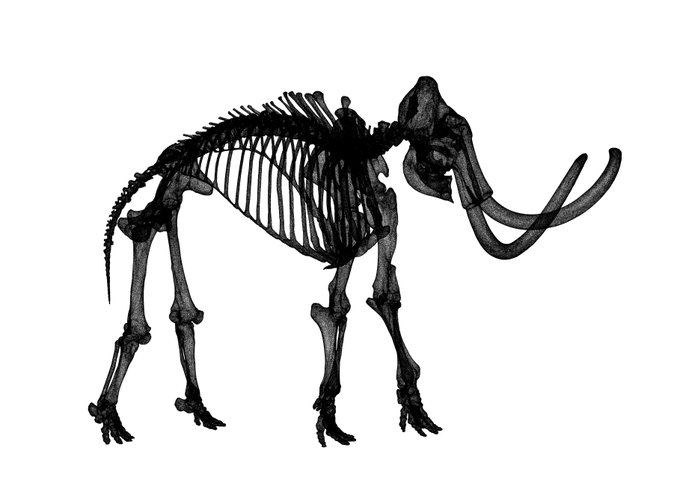
This is a nice way to get some sense of what UMAP is doing at least for low dimensional data.
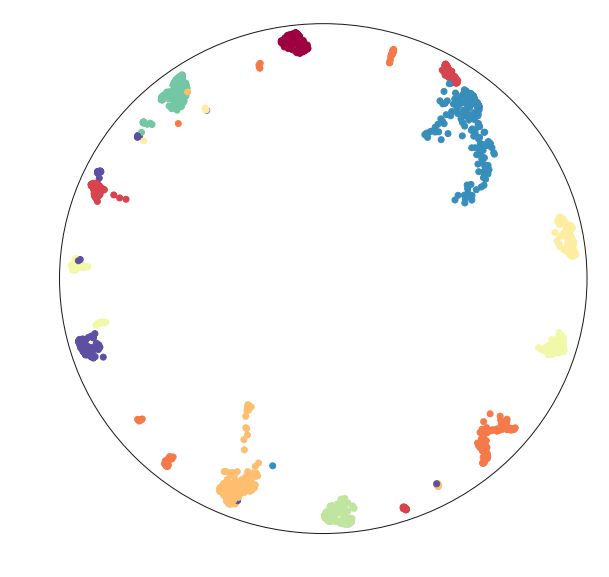
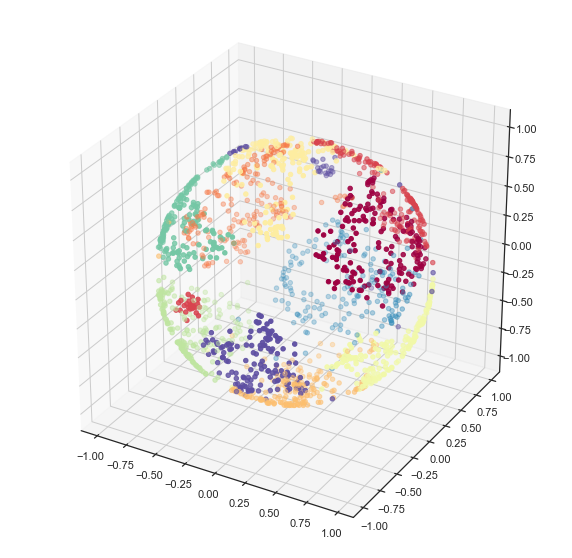
2D UMAP of a 3D woolly mammoth, to build intuitions about how features are preserved in dimensionality reduction. Wonderful 3D scan from the people at
@3D_Digi_Si
.
5
27
76
0
26
87
I really want to emphasize how amazing
@numba_jit
is. Pynndescent is pure python code relying on numba for acceleration. It is performance competitive with *highly optimized* C++ code. I still can't actually believe how incredibly well numba works!
3
20
78
Delivery is apparently a little slower to Canada, but I finally got my copy of
@math3ma
's book! Certainly worth the wait...
2
5
79
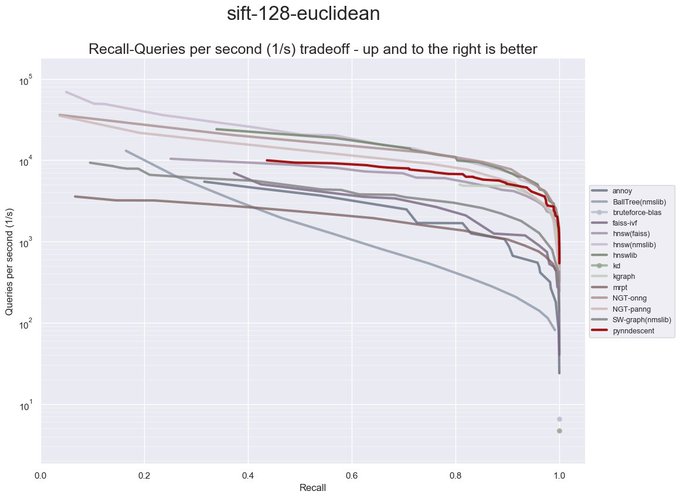
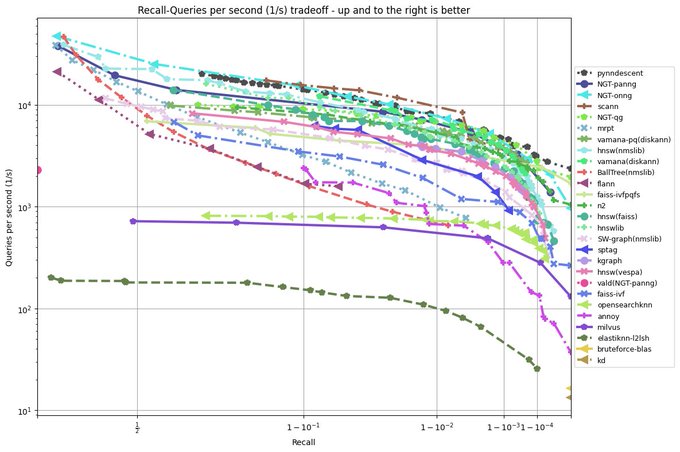
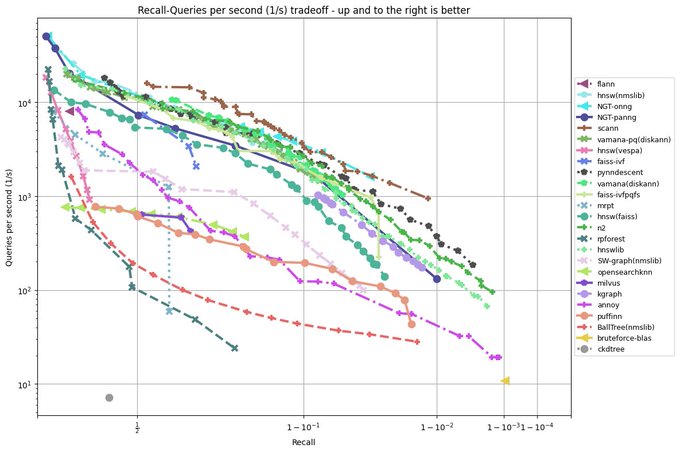
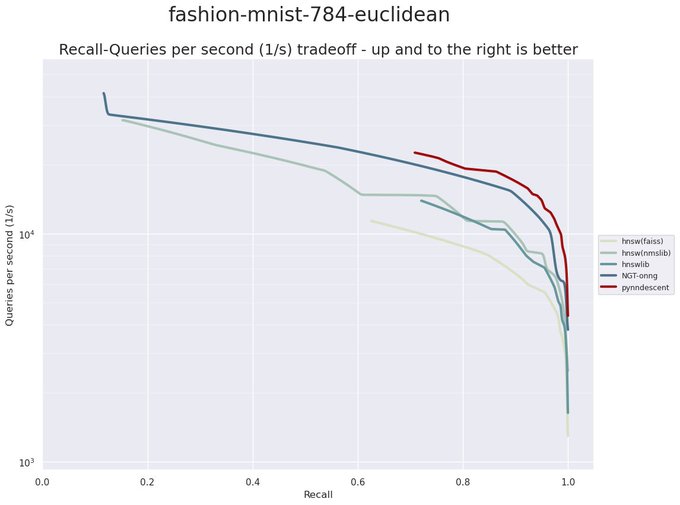
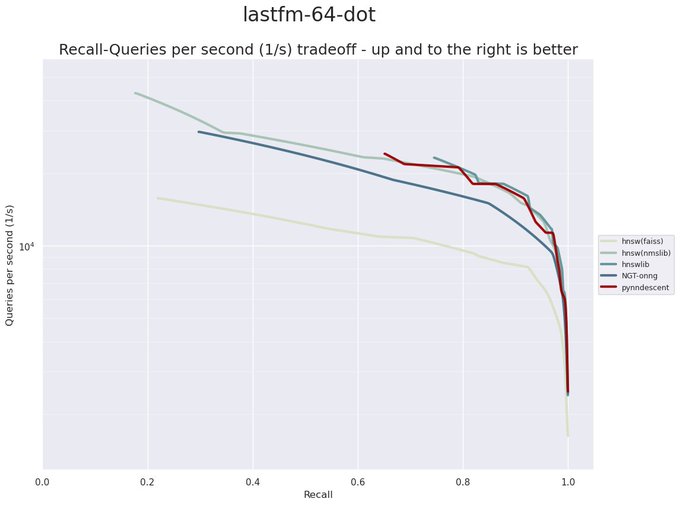
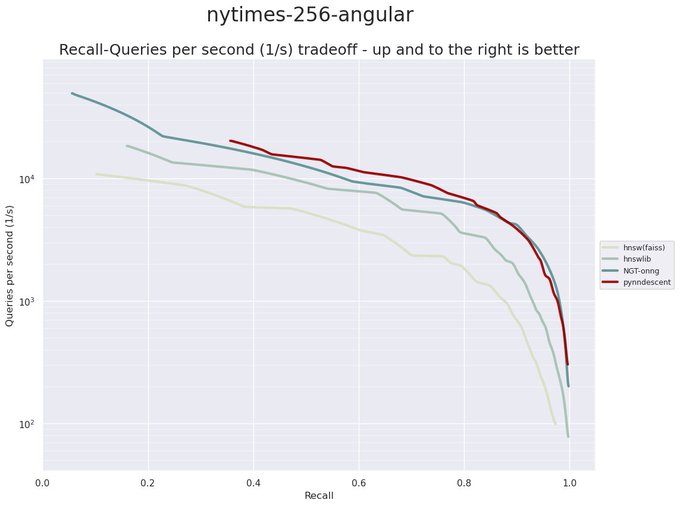
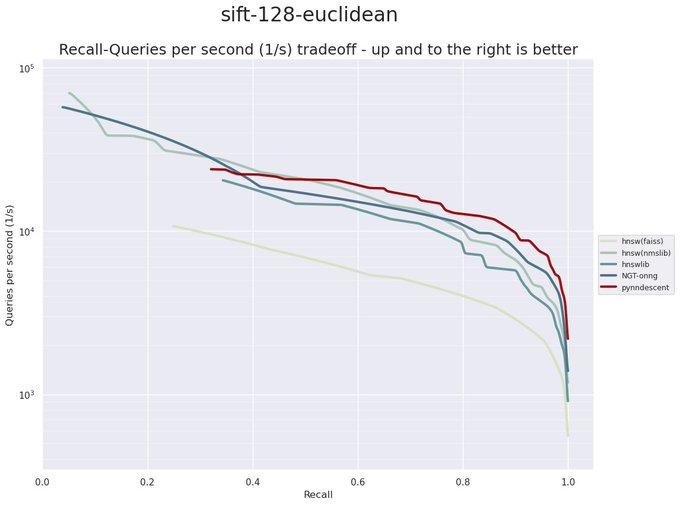
I have been revisiting pynndescent recently, and with help from the
@numba_jit
team I managed to get some significant performance gains. Preliminary tests on
@fulhack
's ann-benchmarks is looking very promising. Hopefully I'll have a new 0.5 release with these changes out soon.
2
17
74
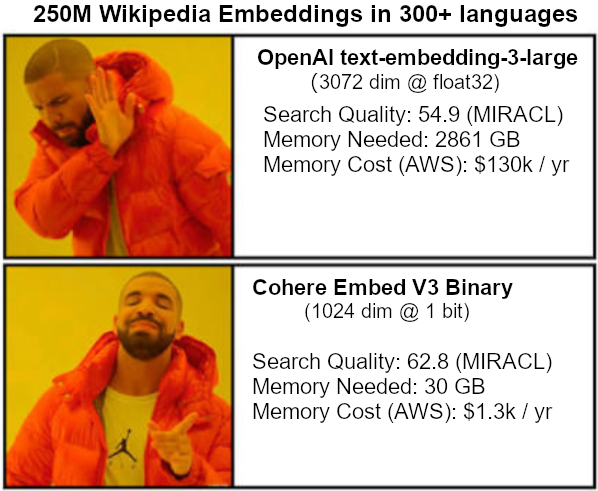
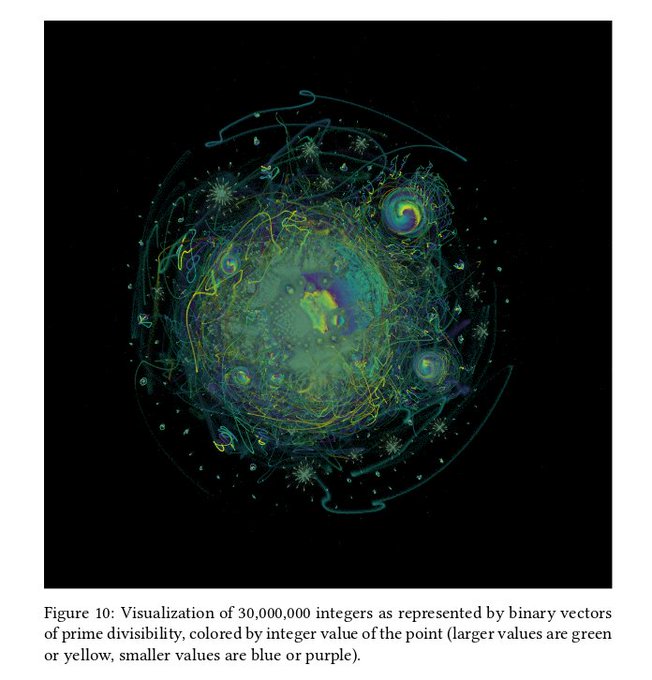
I just added support for these binary embedding vectors to pynndescent. Using them directly with UMAP should be possible very soon...
4
8
72
A paper in
@JOSS_TheOJ
for the UMAP software implementation is now published: .
Thanks to the editors (
@arokem
) and reviewers (
@TerryTangYuan
) for providing such a smooth process for publication.
1
23
65
@DrPattiJones
PCA provides a global linear projection onto the hyperplane defined by the directions of global maximal variance in your data. UMAP attempts to stitch together many local views of the data accounting for local variance, into an intermediate structure, then represent that in low D
3
3
60
Inspired by the t-SNE animation from
@ChaseClarkatUIC
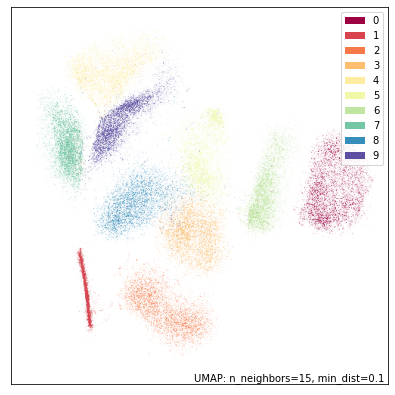
I decided to try something similar for UMAP. Here is an animation for varying values of the n_neighbors parameter. Increasing values give more weight to global structure over local structure.
2
20
60
@ch402
@SuhnyllaKler
@AnthropicAI
An example of current work: is linear optimal transport applied to word vectors a decent sentence/document embedding model? It turns out yes, yes it is.
There's still a long way to go to scale and benchmark on larger datasets, but it's promising.
4
11
56
A new minor release of umap-learn adds some very useful features:
- Updating ParametricUMAP to Keras3 (kindly provided submitted by
@fchollet
);
- Initial support for binary embedding vectors with metric="bit_hamming" and metric="bit_jaccard".
1
3
57
I'll be giving a talk on PyNNDescent, a library for approximate nearest neighbour search, at
#SciPy2021
on Friday.
0
10
54
Code from my lightning talk: ensemble topic modelling in Python with pLSA for fast stable topic modelling with the enstop package:
#SciPy2019
1
17
51
It's well worth reading the paper on FIt-SNE -- useful techniques and fun math.
@F_Vaggi
@leland_mcinnes
FIt-SNE uses an O(N) interpolation scheme to accelerate the computation of the gradient at each step. More details are available in the preprint () or some notes I wrote ()
1
1
20
1
9
48
I belatedly got to experimenting with FIt-SNE from
@GCLinderman
. It's very impressive and very fast -- definitely the implementation you should be using if you want to use t-SNE for visualization.
1
11
48
Good news for
#rstats
users looking for dimension reduction: An R package wrapping UMAP: ; and an independent implementation of UMAP in R: !
0
25
46
The ambient coordinates of your data (coming from features) need not be related to the intrinsic notion of distance internal to the data itself. An idea worth wrapping your head around.
4
14
42
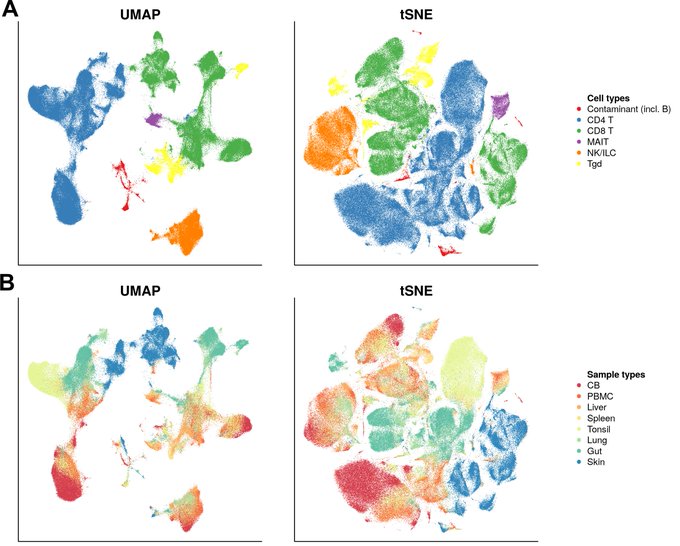
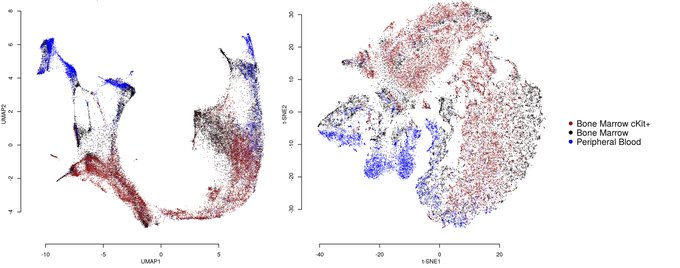
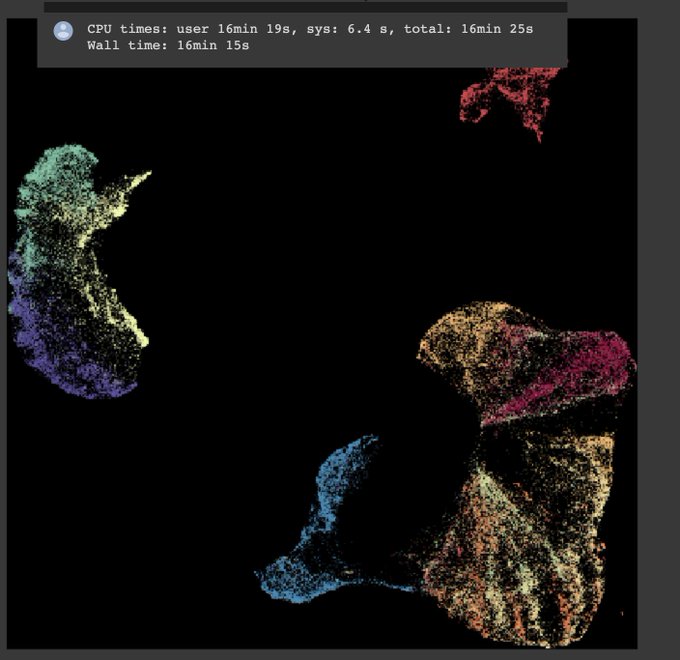
Really interesting to see UMAP on real-world data!
Checkout Etienne Becht's bioRxiv preprint that compares UMAP with t-SNE for visualizing CyTOF and scRNAseq data. Many advantages of UMAP over t-SNE for high dimensional single-cell data!
@leland_mcinnes
11
109
227
2
10
43
This was a fantastic series of of posts! If you want a well written intro to some of the ideas in topological data analysis this is a great place to start.
@asemic_horizon
@scikit_tda
@leland_mcinnes
I wrote a series of posts leading up to some TDA (see "Topology" section here: ) And then a few posts in the TDA family before I lost steam (see Computational Topology section of )
1
4
36
1
13
42
Many thanks to
@datametrician
@cjnolet
and
@rapidsai
for making this possible -- definitely some amazing performance available for UMAP on GPU!
Reproduced the
#UMAP
on
#RAPIDS
example by
@ceshine_en
() on Colab (with help from ). Seeing 60X speedup on Colab
@leland_mcinnes
@rapidsai
@keithjkraus
@datametrician
@rodaramburu
see Colab Gist
0
13
44
1
18
41
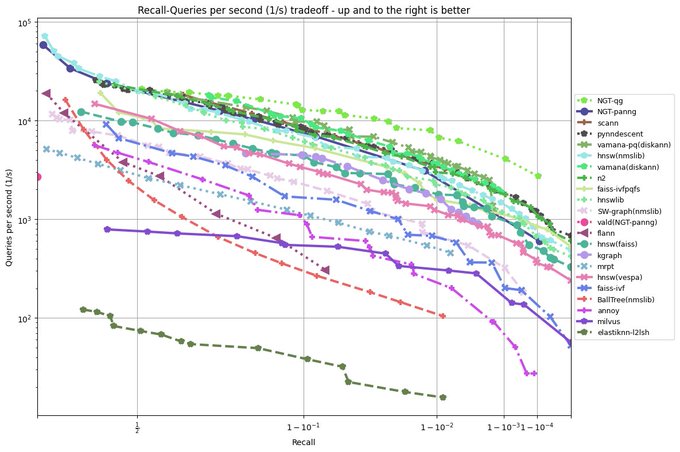
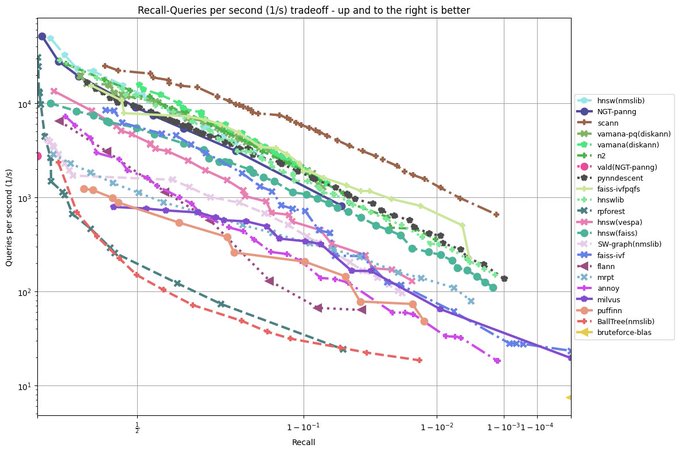
It is a huge testament to the power of
@numba_jit
that a pure python library like PyNNDescent can be performance competitive with C++ libraries from Google (ScaNN), Microsoft (DiskANN), and Facebook (FAISS) among others.
Many, many thanks to the whole
@numba_jit
team!
3
6
40
@EmilyTWinn13
@SC_Griffith
After the flood Noah is checking up on the animals. They're all breeding well, except for a pair of snakes. Noah gets a little worried and follows them. Eventually they find a fallen tree, and suddenly ... lots of baby snakes. It turns out that adders need logs to multiply.
1
4
39
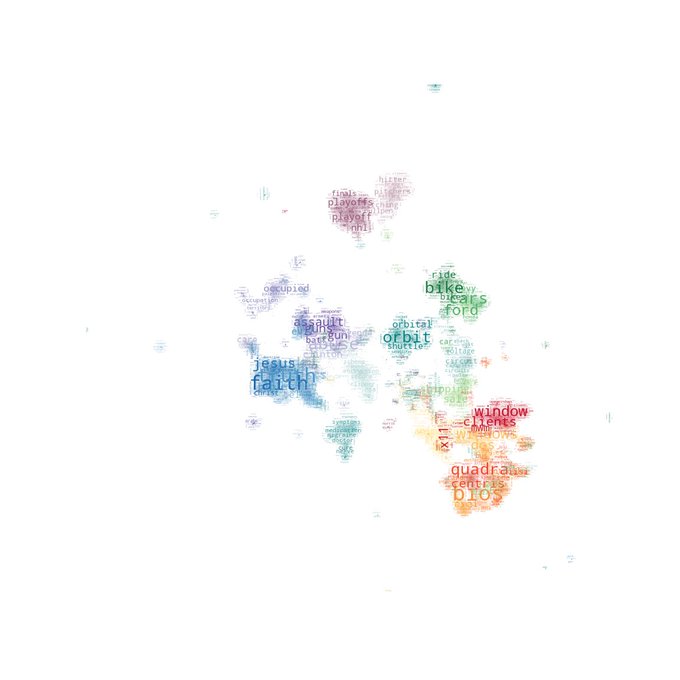
@rctatman
Here's a plan we use: Take the term-frequency matrix, remove the "expected" frequency (by subtracting, or using the column marginal as a noise model), UMAP with hellinger distance, and HDBSCAN for clustering. Still fine tuning the process, but has been very powerful so far.
3
2
39
An amazing introduction to UMAP and its parameters. This is for UMAP what the Distill article was for t-SNE. Great work from
@_coenen
and
@adamrpearce
as always!
1
19
39
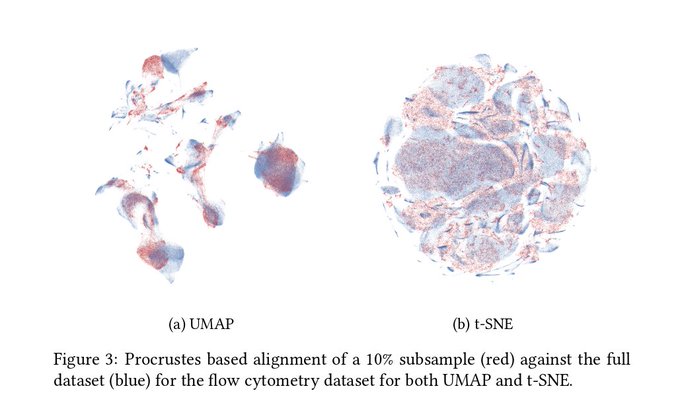
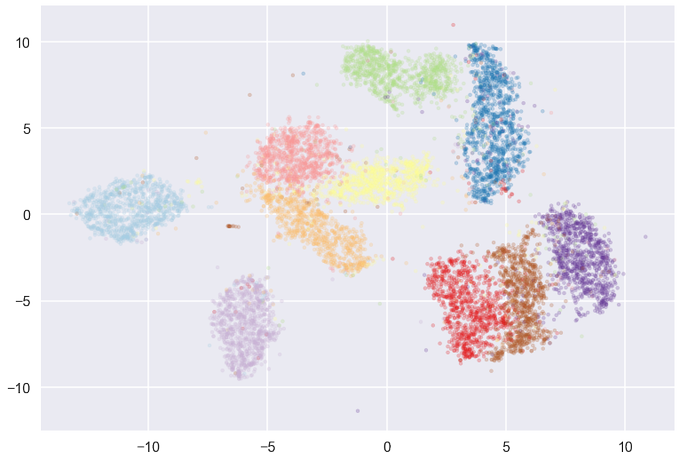
@michaelhoffman
Many of the t-SNE (and UMAP) plots I see suffer from potential over-plotting issues. This is particularly dangerous if you are trying to eyeball cluster purity. Using such plots as a starting point for further analysis rather than an endpoint is critical.
4
10
38
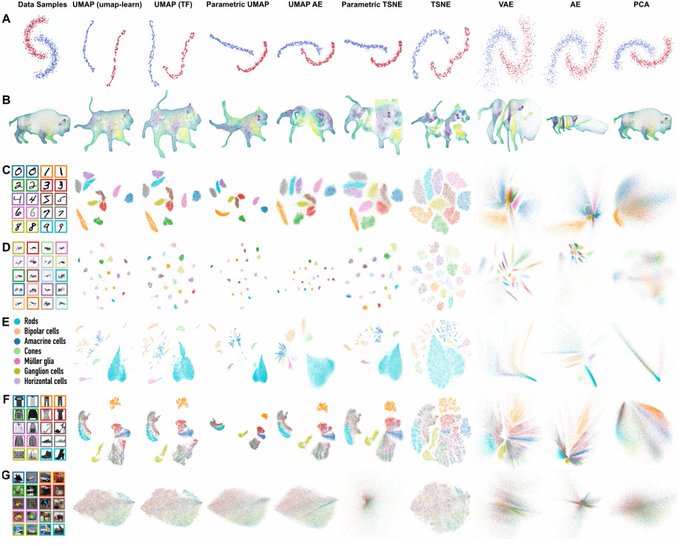
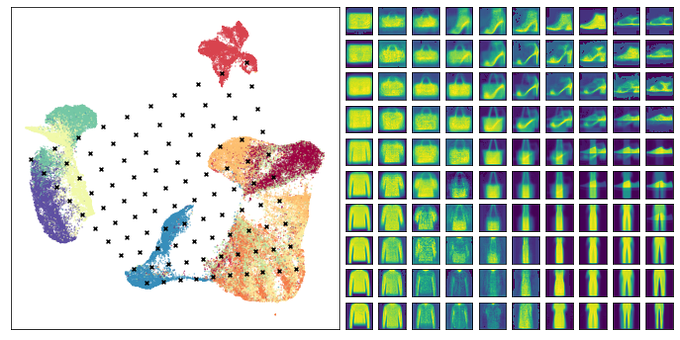
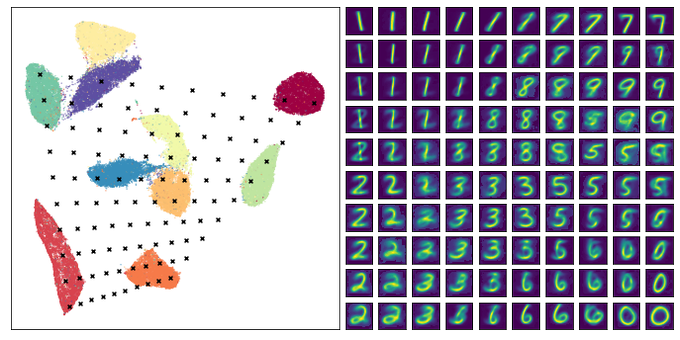
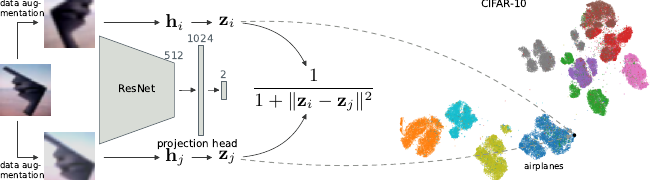
This is a fascinating paper -- using a contrastive approach on augmentations of images to learn a low dimensional representation they generate truly impressive results for image datasets!
Ever wondered what image datasets look like if they could be visualized? We have developed a new algorithm for visualization based on contrastive learning. Joint work with
@hippopedoid
and
@CellTypist
. The full details are available as a preprint 🧵/16
4
66
264
1
3
37
I've started telling people "Look at your data, because whatever you think you know about the data is almost certainly wrong". I'm not sure it works any better, but at least I warned them...
2
11
36
Here's a really great interactive article using UMAP to explore and compare large deep neural networks by
@mwli16
and
@scheidegger
:
0
9
35
I will be co-chairing the machine learning track at SciPy this year. Submissions are open, so if you have a machine learning project in python consider submitting. This is a great opportunity to share your work with a wide audience.
@SciPyConf
0
6
36
Getting close to finishing version 0.3 of UMAP, including some useful new features. Ideally it'll come at just before or at
@SciPyConf
this year.
2
4
34